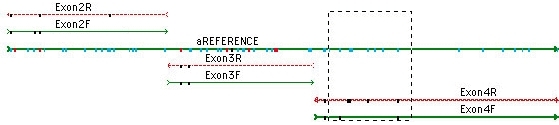
All sequences analyzed in this study were deposited at GenBank and are listed in Table 2. The newly generated sequences were deposited at GenBank ( (accessed on 8 December 2022)). The PCR products were purified and sequenced in Bioengineering (Shanghai) Co., Ltd., China, with the same primers. The PCR procedure for nrLSU was as follows: initial denaturation at 94 ☌ for 4 min, followed by 35 cycles at 94 ☌ for 1 min, 53 ☌ for 1 min, 72 ☌ for 1 min, and a final extension of 72 ☌ for 10 min. The PCR procedure for ITS (including 5.8 S) was as follows: initial denaturation at 94 ☌ for 4 min, followed by 35 cycles at 94 ☌ for 35 s, 54 ☌ for 35 s, and 72 ☌ for 45 s, and a final extension of 72 ☌ for 10 min. LR0R (5′-ACCCGCTGAACTTAAGC-3′) and LR5 (5′-ATCCTGAGGGAAACTTC-3′) were used for the large subunit of the nuclear ribosomal RNA gene (nrLSU). Two molecular markers were investigated, i.e., ITS1F (3′-CTTGGTCATTTAGAGGAAGTAA-5′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′), which were used as primers for the internal transcribed spacer (ITS) (White et al. The 30 μL PCR reaction system is shown in Table 1. Genomic DNA was extracted from 0.1 to 0.2 mg of dried specimen using a NuClean Plant Genomic DNA kit (CWBIO, Beijing, China) and preserved at −20 ☌. The range ‘c–d’ means the minimum to maximum length, ‘e–f’ and‘g–h’ means the minimum to maximum width. The range ‘a–b’ means the minimum to the maximum of the diameter. 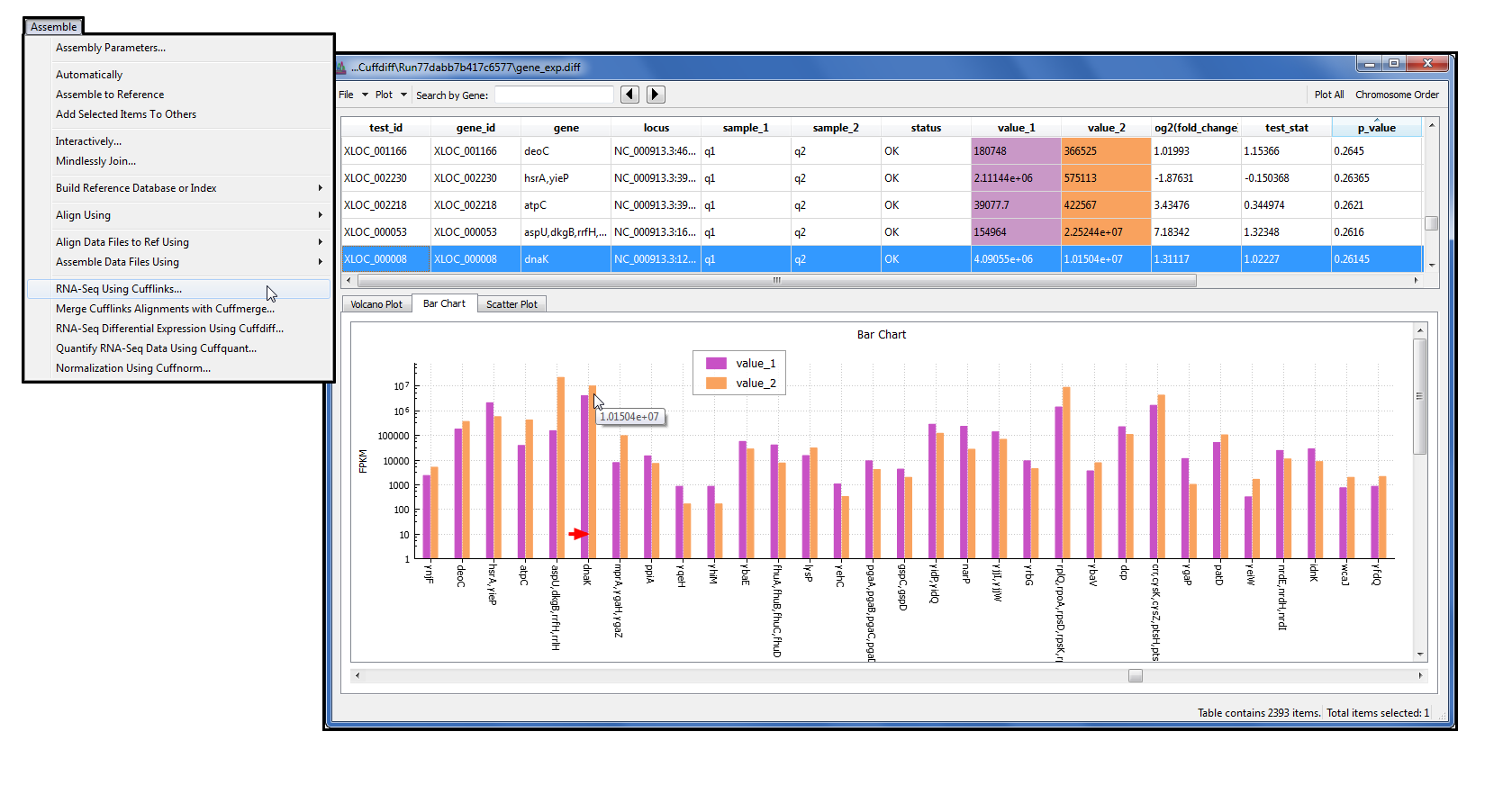 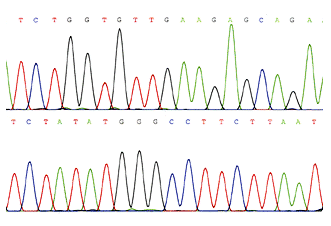
The abbreviation represents n basidiospores measured from m basidiomata of p collections. The dimensions of capillitial hyphae are given using a notation in the form ‘g–h’. The dimensions of basidium are given using a notation in the form ‘c–d × e–f’. The dimensions of basidiospores are given using a notation in the form ‘a–b’. Data of the sections (basidiospores, basidia, capillitial hyphae, and exoperidium) were obtained from dried specimens, which were rehydrated in 5% KOH or stained in Congo red when necessary, and the light microscope (Olympus BX50) was used for the examination of microscopic structures with a high-resolution oil objective lens (1000×). Dried specimens were used to observe microscopic features. Micromorphological studies were carried out using a light microscope and scanning electron microscope. The color description of the basidiomata was based on Kornerup and Wanscher (1978). Macromorphological descriptions were based on fresh specimens, which were photographed in the field with notes and laboratory supplemental measurements. They are highly divergent due to the different relative value that each author gave to particular morphological features, so it is difficult to group using traditional categorization methods alone, calling for a need to combine the support of molecular data. Because the Geastrum genus has few classification features, its subgenera are mainly distinguished according to morphological features. In addition, Lloyd (1902) proposed that depending on whether the exoperidium is hygroscopic, this factor can be used as a basis for categorization and Geastrum can be divided into two sections, viz., Sect. 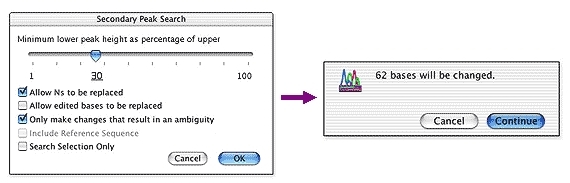
Dörfelt (1985) divided the Geastrum genus into four subgenera, viz., Subg. Ponce (1968) divided the Geastrum into one subgenus and six section, viz., Subg. Section Basimyceliata was further revised by Dissing and Lange (1962), based on the integrity of the endoperidium, and subdivided into three subsections and eight species. Basimyceliata this was later endorsed by Sunhede (1989). Staněk (1958) proposed dividing Geastrum into two sections based on differences between the peristomes and whether an encrustation of debris is present, viz., Sect. This was later endorsed by Hollós (1903). Then De Toni (1887) proposed dividing the Geastrum genus into seven sections based on the morphological characteristics of the peristome, stalk, and exoperidium, viz., Sect. Geastrum was first identified by Persoon (1794).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
